|
We think that this is a really cool paper, describing a method to use PacBio sequencing to look at microbial eukaryotic diversity.
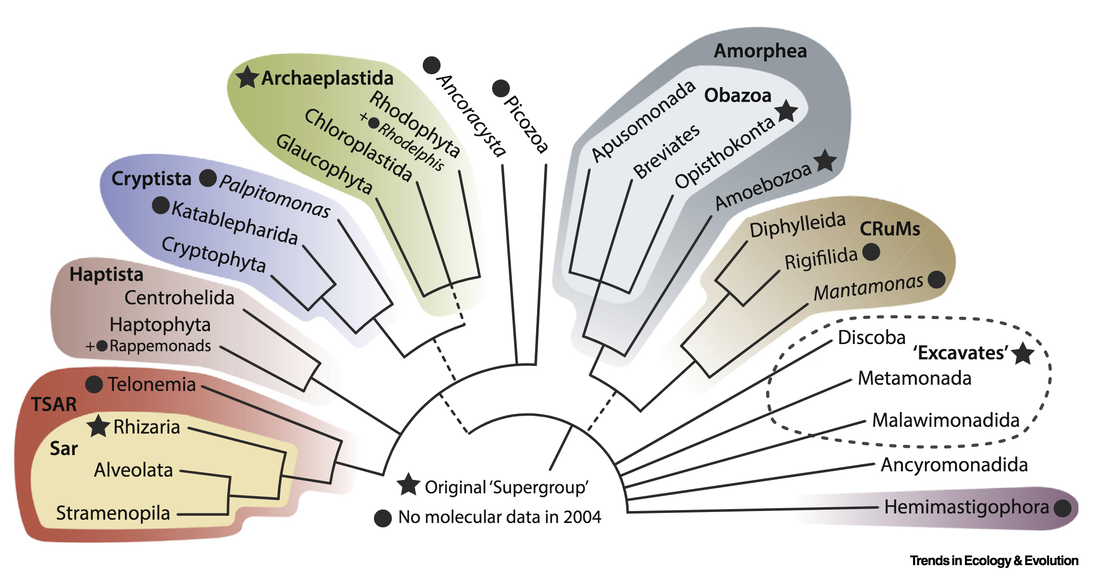
See for yourself here: onlinelibrary.wiley.com/doi/10.1111/1755-0998.13117 Together with Alastair Simpson and Andrew Roger (Dalhousie University), and Matt Brown (Mississippi State University), Fabien has just published in Trends in Ecology and Evolution a review paper on the tree of eukaryotes. Check it out here: www.cell.com/trends/ecology-evolution/fulltext/S0169-5347(19)30257-5
Postdoctoral Fellow in Marine Microbiology with Emphasis on Endosymbiosis
This is a joint position with the lab of Dr. Rachel Foster at Stockholm University, to work on a poorly known, yet exciting endosymbiosis in the protistan genus Meringosphaera. Deadline for applying is November 5, 2019. Full description and how to apply: www.su.se/english/about/working-at-su/jobs?rmpage=job&rmjob=10173&rmlang=UK Nikiforos has just joined the lab for a research practice has part of his Master degree at Uppsala University. He will work on assessing picozoan diversity in metagenomic datasets. Most welcome!
Together with Patrick Keeling from UBC, Fabien just published a review paper discussing the current view on the eukaryotic tree of life (eToL). Among other topics, the fitting of environmental diversity in the tree and its increasingly lopsided appearance are discussed. Check it out: https://doi.org/10.1016/j.cub.2019.07.031
Long metabarcoding of the eukaryotic rDNA operon to phylogenetically and taxonomically resolve environmental diversity. This is a paper where we describe the use of pacbio to sequence eDNA, and apply a novel approach to resolve the environmental diversity of microbial eukaryotes (protists) using phylogenetics.
Link to the pre-print: https://www.biorxiv.org/content/10.1101/627828v1 We welcome Elena Coulier in the team. Elena is from the Université Clermont Auvergne in France and will be with us for a 3 months internship as part of her studies.
Fabien recently became Associate Editor for Molecular Phylogenetics and Evolution, handling protist papers. The goal is to increase the protist-focused paper in this journal, so please send your manuscript along!
Boum boum, 2 new papers from the lab were recently published:
We recently said bye to Jürgen, who went back to Germany after a successful 2 years in the lab. Sad, but we welcomed a new member last week: Anders Alfjorden, who is doing a PhD on various aspects of Ascetosporean parasites.
|